Functional Coupling between Transcription and RNA Processing
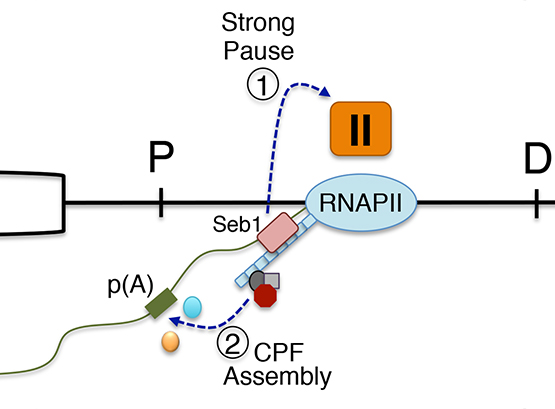
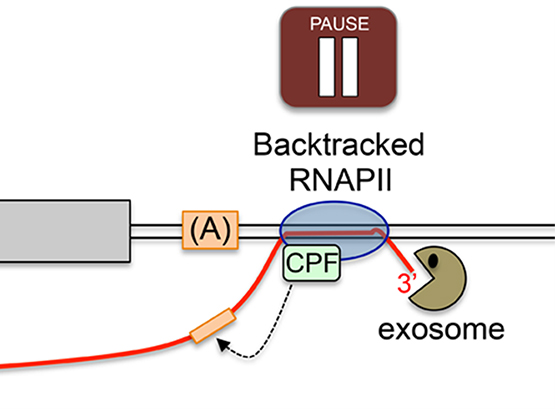
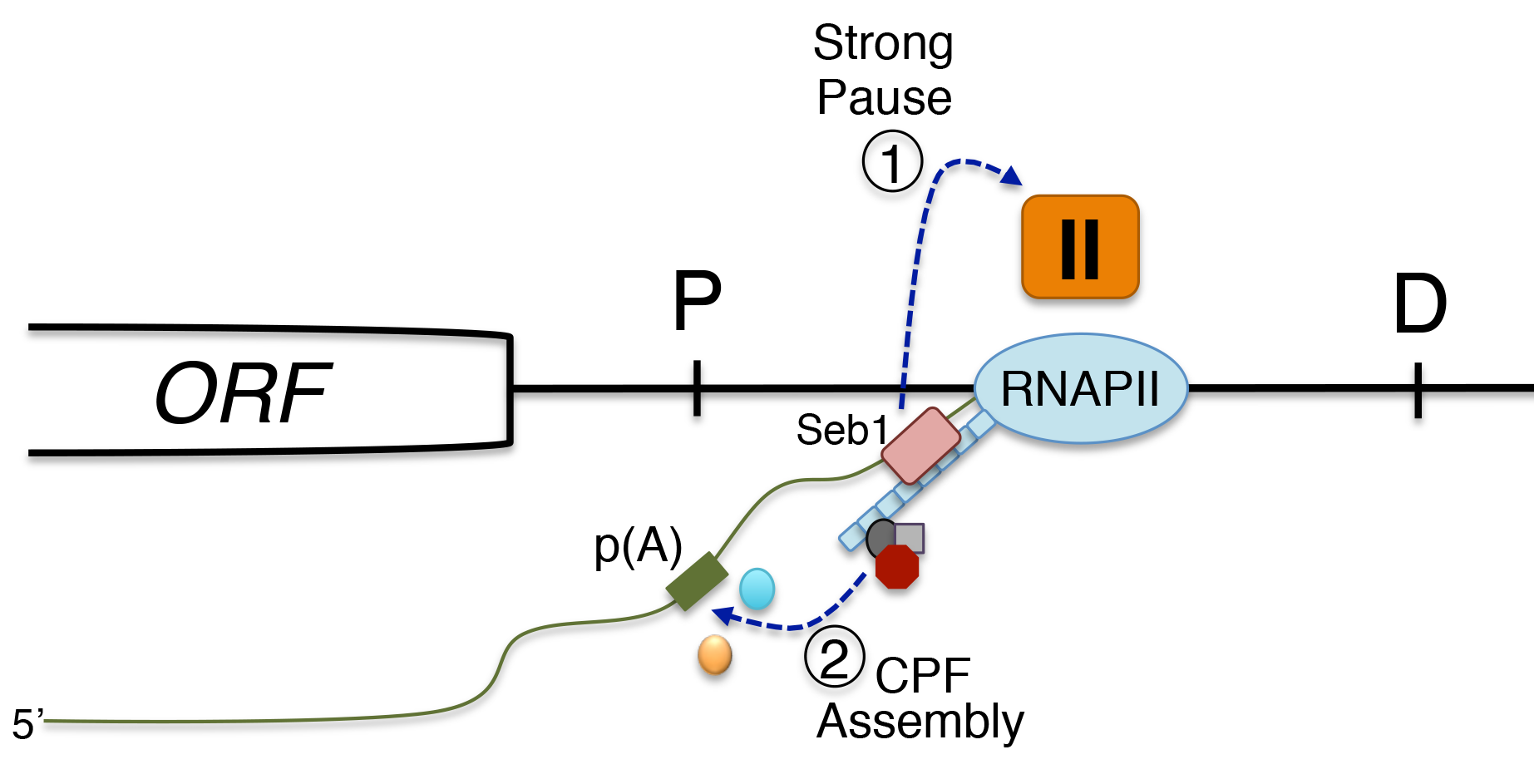
We are highly interested in understanding how various steps of RNA processing are intimately coupled to transcription by RNA polymerases. In the past few years, we have discovered that the conserved fission yeast protein Seb1 plays a critical role in polyadenylation site selection. Seb1 is recruited to transcription units via the C-terminal domain (CTD) of the largest subunit of the RNA polymerase II complex where Seb1 positively contributes to transcription pausing, thereby promoting polyadenylation site selection.
We have also used genome editing by CRISPR/Cas9 in yeast to introduce a substitution in Rbp1 that reduces the transcription elongation rate and showed that this RNAPII mutations causes widespread changes in alternative polyadenylation (APA).
Feature publications:
Yague-Sanz, C., Y. Vanrobaeys, R. Fernandez, M. Duval, M. Larochelle, J. Beaudoin, J. Berro, S. Labbe, P. E. Jacques and F. Bachand (2020). “Nutrient-dependent control of RNA polymerase II elongation rate regulates specific gene expression programs by alternative polyadenylation.” Genes Dev 34: 883-897.
Larochelle, M., M. A. Robert, J. N. Hebert, X. Liu, D. Matteau, S. Rodrigue, B. Tian, P. E. Jacques and F. Bachand (2018). “Common mechanism of transcription termination at coding and noncoding RNA genes in fission yeast.” Nat Commun 9: 4364.
Lemay, J. F., S. Marguerat, M. Larochelle, X. Liu, R. van Nues, J. Hunyadkurti, M. Hoque, B. Tian, S. Granneman, J. Bahler and F. Bachand (2016). “The Nrd1-like protein Seb1 coordinates cotranscriptional 3′ end processing and polyadenylation site selection.” Genes Dev 30: 1558-1572.
BROAD RESEARCH PROGRAM
DETAILS ON FALLOWING PROJECTS
Polyadenylation-dependent Gene Regulation
Oculopharyngeal muscular dystrophy (OPMD) is an inherited disease caused by mutations in the gene encoding PABPN1 (Poly(A)-Binding Protein Nuclear 1).
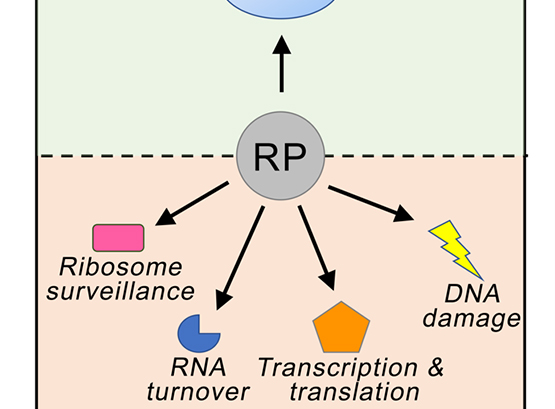
Extra-ribosomal functions of Ribosomal Proteins
The general view of ribosomal proteins (RPs) is one of generic cellular proteins that function exclusively to maintain the integrity of the ribosome.
Functional Coupling between Transcription and RNA Processing
We are highly interested in understanding how various steps of RNA processing are intimately coupled to transcription by RNA polymerases.
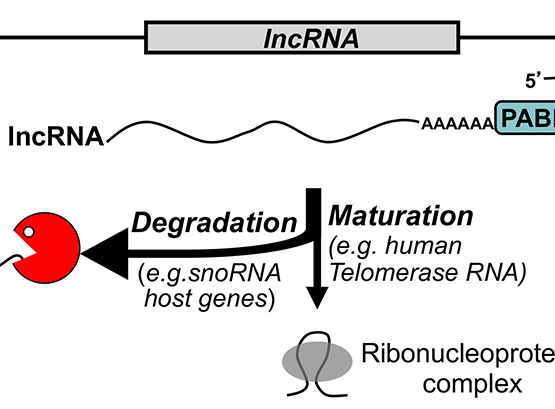
Nuclear RNA surveillance
Recent studies indicate that transcription is pervasive in eukaryotes, producing a complex network of noncoding RNAs (ncRNAs) that exist stably in cells or are rapidly degraded.