Nuclear RNA surveillance
Recent studies indicate that transcription is pervasive in eukaryotes, producing a complex network of noncoding RNAs (ncRNAs) that exist stably in cells or are rapidly degraded. To date, however, little is known about the pathways and mechanisms that promote RNA surveillance and quality control.
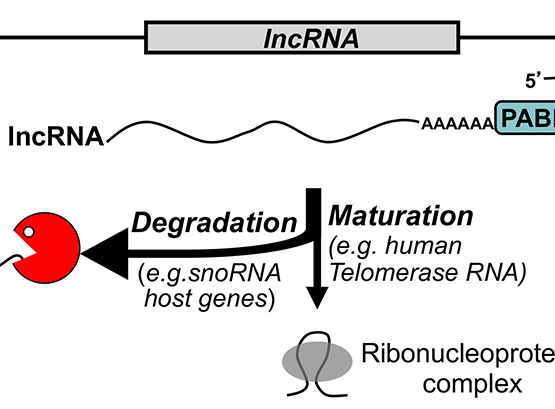
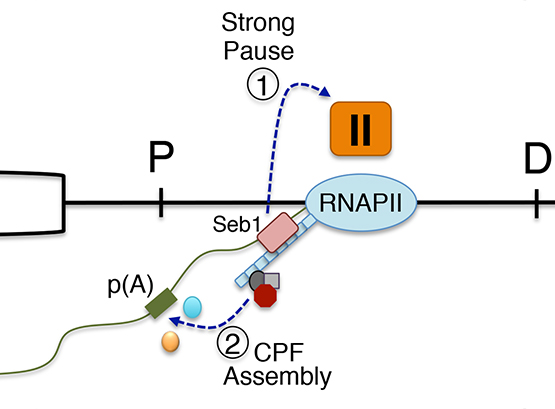
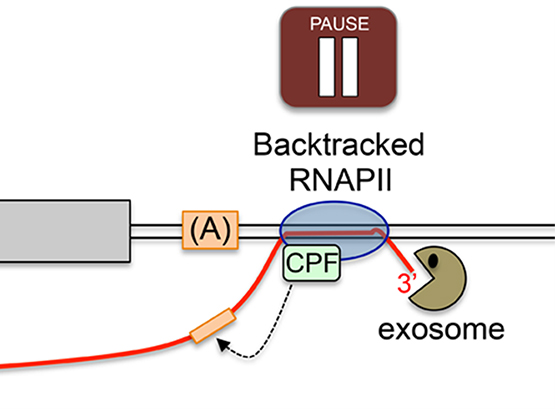
Our work on the exosome complex of 3’-5’ exonucleases has uncovered a novel mechanism of transcription termination that prevents the accumulation of aberrant read-through RNAs and transcriptional interference at neighboring genes. We also made a major breakthrough into the understanding of how polyadenylation site selection is intimately coordinated with RNA quality control.
Feature publications:
Bitton, D. A., S. R. Atkinson, C. Rallis, G. C. Smith, D. A. Ellis, Y. Y. Chen, M. Malecki, S. Codlin, J. F. Lemay, C. Cotobal, F. Bachand, S. Marguerat, J. Mata and J. Bahler (2015). “Widespread exon skipping triggers degradation by nuclear RNA surveillance in fission yeast.” Genome Res 25: 884-896.
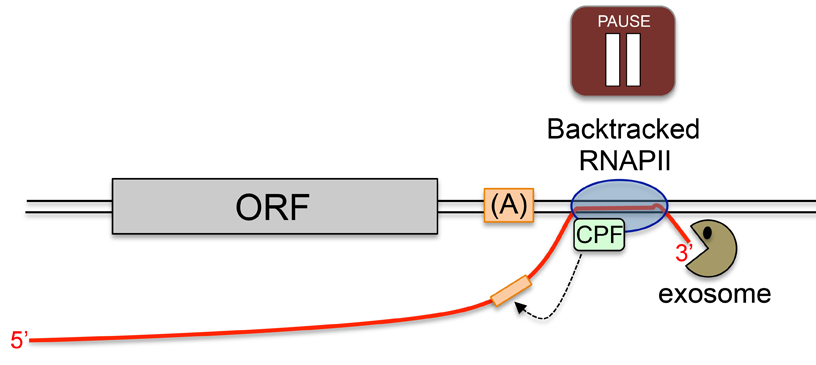
Lemay, J. F., M. Larochelle, S. Marguerat, S. Atkinson, J. Bahler and F. Bachand (2014). “The RNA exosome promotes transcription termination of backtracked RNA polymerase II.” Nat Struct Mol Biol 21: 919-926.
Larochelle, M., J. F. Lemay and F. Bachand (2012). “The THO complex cooperates with the nuclear RNA surveillance machinery to control small nucleolar RNA expression.” Nucleic Acids Res 40: 10240-10253.
BROAD RESEARCH PROGRAM
DETAILS ON FALLOWING PROJECTS
Polyadenylation-dependent Gene Regulation
Oculopharyngeal muscular dystrophy (OPMD) is an inherited disease caused by mutations in the gene encoding PABPN1 (Poly(A)-Binding Protein Nuclear 1).
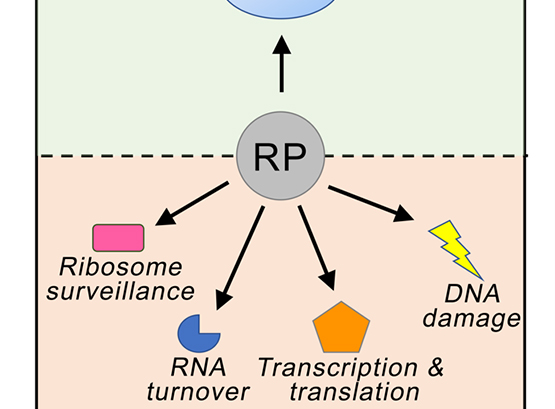
Extra-ribosomal functions of Ribosomal Proteins
The general view of ribosomal proteins (RPs) is one of generic cellular proteins that function exclusively to maintain the integrity of the ribosome.
Functional Coupling between Transcription and RNA Processing
We are highly interested in understanding how various steps of RNA processing are intimately coupled to transcription by RNA polymerases.
Nuclear RNA surveillance
Recent studies indicate that transcription is pervasive in eukaryotes, producing a complex network of noncoding RNAs (ncRNAs) that exist stably in cells or are rapidly degraded.